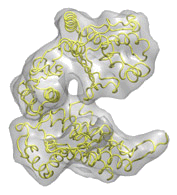
| We develop and apply new hybrid methods to combine multiresolution structural information in collaboration with several electron microscopy (EM) labs. Multiresolution fitting is the standard way to interpret the macromolecular EM maps by employing available atomic structural components. This process is akin to assembling a jigsaw puzzle, where a lower-resolution 3D EM density map of a macromolecular complex acts as a fuzzy frame to guide the integration of interlocking atomic-resolution pieces. When complete, this jigsaw puzzle allows us to characterize different functional states of macromolecules in solution at near-atomic detail. We have been involved in developing several approaches to solve this puzzle. Firstly, we contributed to the pioneering EM modeling package, Situs. Additionally, complementary tools for rigid-body fitting (ADP_EM) and flexible fitting (iMODfit, which are fully available on our site, have been developed and successfully applied in various real-world fitting scenarios. |
 |